Overview
SIAMCAT is a pipeline for Statistical Inference of Associations between Microbial Communities And host phenoTypes. A primary goal of analyzing microbiome data is to determine changes in community composition that are associated with environmental factors. In particular, linking human microbiome composition to host phenotypes such as diseases has become an area of intense research. For this, robust statistical modeling and biomarker extraction toolkits are crucially needed. SIAMCAT provides a full pipeline supporting data preprocessing, statistical association testing, statistical modeling (LASSO logistic regression) including tools for evaluation and interpretation of these models (such as cross validation, parameter selection, ROC analysis and diagnostic model plots).
SIAMCAT is developed in the Zeller group and is part of the suite of computational microbiome analysis tools hosted at EMBL.
Starting with SIAMCAT
Installation
In order to start with SIAMCAT, you need to install it from Bioconductor:
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("SIAMCAT")Alternatively, you can install the current development version via devtools:
require("devtools")
devtools::install_github(repo = 'zellerlab/siamcat')Quick start
There are a few manuals that will kick-start you and help you analyse your data with SIAMCAT. You can find links to those on the Bioconductor website of SIAMCAT or you can type into R:
browseVignettes("SIAMCAT")
# Please Note:
# `browseVignettes` only works if `SIAMCAT` has been installed via BioconductorFeedback and Contact
If you have any question about SIAMCAT, if you run into any issue, or if you would like to make a feature request, please: - create an issue in this repository or - mail Georg Zeller or - ask at the SIAMCAT support group
If you have a more general question (that could be useful to several other users), please do not hesitate to post it on a dedicated forums such as Stackoverflow or Biostars. If you let us know about the question, we will answer it swiftly.
Please consider giving us feedback. (This feedback is useful for us to justify the funding we get for developing and maintaining this package.)
License
SIAMCAT is distributed under the GPL-3 license.
Citation
If you use SIAMCAT, please cite us by using
citation("SIAMCAT")or by
Wirbel J, Zych K, Essex M, Karcher N, Kartal E, Salazar G, Bork P, Sunagawa S, Zeller G Microbiome meta-analysis and cross-disease comparison enabled by the SIAMCAT machine learning toolbox Genome Biol 22, 93 (2021) https://doi.org/10.1186/s13059-021-02306-1
In this publication, we analyzed a large set of case-control microbiome datasets. The metadata and taxonomic profiles of these studies are available through a Zenodo repository: .
Examples of primary package output
To give you a small preview about the primary package output, here are some example plots taking from the main SIAMCAT vignette.
In this vignette, we use an example dataset which is also included in the SIAMCAT package. The dataset is taken from the publication of Zeller et al, which demonstrated the potential of microbial species in fecal samples to distinguish patients with colorectal cancer (CRC) from healthy controls.
Association testing
The result of the check.associations function is an association plot. For significantly associated microbial features, the plot shows: - the abundances of the features across the two different classes (CRC vs. controls) - the significance of the enrichment calculated by a Wilcoxon test (after multiple hypothesis testing correction) - the generalized fold change of each feature - the prevalence shift between the two classes, and - the Area Under the Receiver Operating Characteristics Curve (AU-ROC) as non-parametric effect size measure.
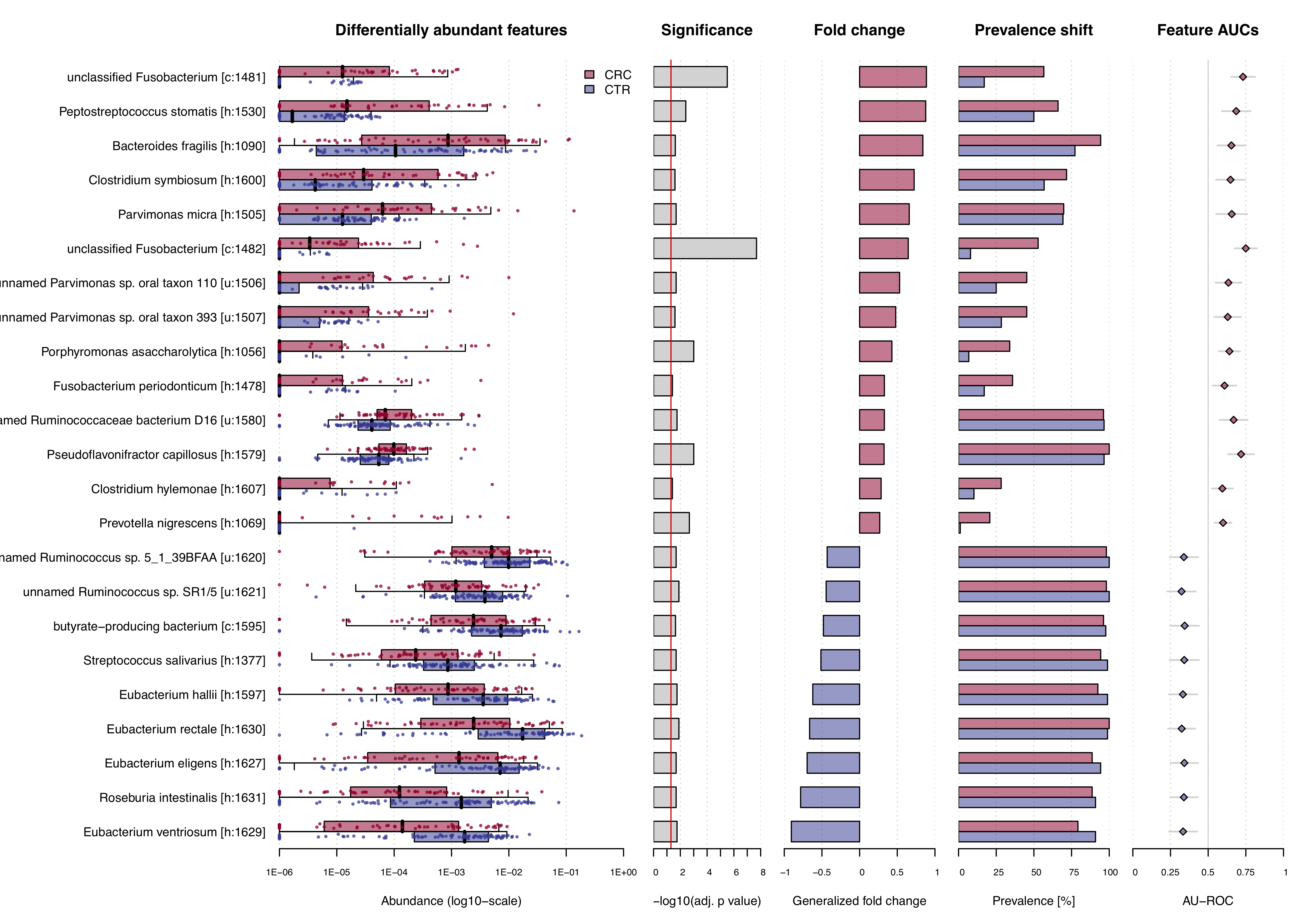
Association testing
Model interpretation plot
After statistical models have been trained to distinguish cancer cases from controls, the models can be investigated by the function model.interpretation.plot. The plots shows: - the median relative feature weight for selected features (barplot on the left) - the robustness of features (i.e. in how many of the models the specific feature has been selected) - the distribution of selected features across samples (central heatmap) - which proportion of the weight of all different models are shown in the plot (boxplot on the right), and - distribution of metadata across samples (heatmap below).
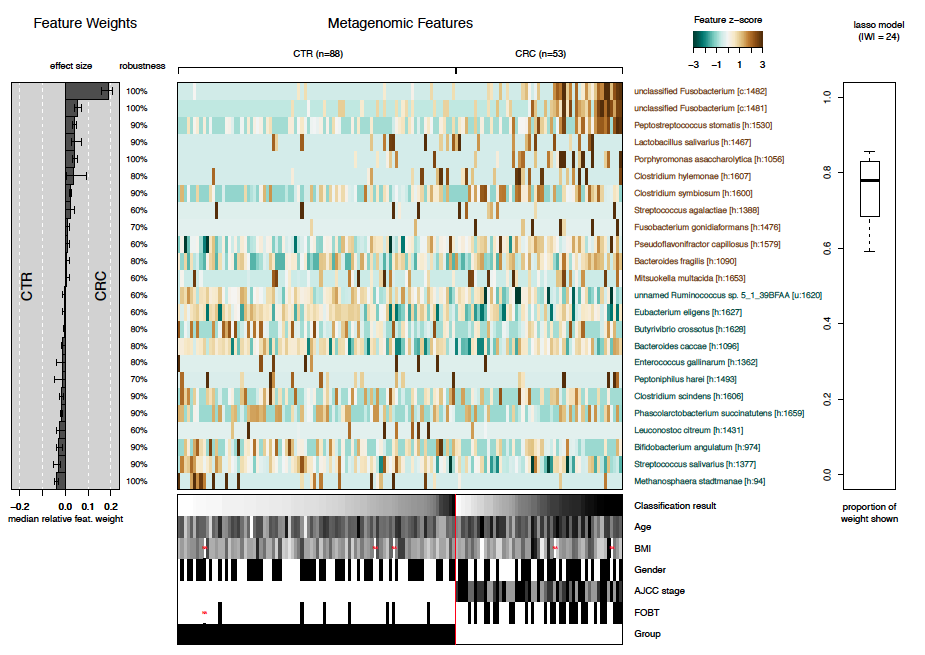
Model interpretation plot
Where SIAMCAT has been used already
Several publications already used SIAMCAT (or previous versions thereof).
Potential of fecal microbiota for early-stage detection of colorectal cancer
Zeller G, Tap J, Voigt AY, Sunagawa S, Kultima JR, Costea PI, Amiot A, Böhm J, Brunetti F, Habermann N, Hercog R, Koch M, Luciani A, Mende DR, Schneider MA, Schrotz-King P, Tournigand C, Tran Van Nhieu J, Yamada T, Zimmermann J, Benes V, Kloor M, Ulrich CM, von Knebel Doeberitz M, Sobhani I, Bork P
Molecular Systems Biology, (2014) 10, 766
>Original Publication that inspiredSIAMCATGut Microbiota Linked to Sexual Preference and HIV Infection
Noguera-Julian M, Rocafort M, Guillén Y, Rivera J, Casadellà M, Nowak P, Hildebrand F, Zeller G, Parera M, Bellido R, Rodríguez C,Carrillo J, Mothe B, Coll J, Bravo I, Estany C, Herrero C, Saz J, Sirera G, Torrela A, Navarro J, Crespo M, Brander C, Negredo E, Blanco J, Guarner F, Calle ML, Bork P, Sönnerborgd A, Clotet B, Paredes R
EBioMedicine 5 (2016) 135-146 >See Figure 5Extensive transmission of microbes along the gastrointestinal tract
Schmidt TSB, Hayward MR, Coelho LP, Li SS, Costea PI, Voigt AY, Wirbel J, Maistrenko OM, Alves RJC, Bergsten E, de Beaufort C, Sobhani I, Heintz-Buschart A, Sunagawa S, Zeller G, Wilmes P, Bork P
eLife, (2019) 8:e42693
> See Figure 3 - figure supplement 1Meta-analysis of fecal metagenomes reveals global microbial signatures that are specific for colorectal cancer
Wirbel J, Pyl PT, Kartal E, Zych K, Kashani A, Milanese A, Fleck JS, Voigt AY, Palleja A, Ponnudurai R, Sunagawa S, Coelho LP, Schrotz-King P, Vogtmann E, Habermann N, Niméus E, Thomas AM, Manghi P, Gandini S, Serrano D, Mizutani S, Shiroma H, Shiba S, Shibata T, Yachida S, Yamada T, Waldron L, Naccarati A, Segata N, Sinha R, Ulrich CM, Brenner H, Arumugam M, Bork P, Zeller G
Nature Medicine, (2019) [Epub ahead of print]
> In this publication,SIAMCATis used extensively for holdout testing
If you used SIAMCAT in your publication, let us know!



